Archive for the ‘Neural Grammar Network’ tag
Back from Conference!
Brief: The BIBM09 conference was the very first conference I have ever attended. I learned a lot from the various speakers and poster sessions–
I thought it was really interesting how the trend is to now study and manipulate large interaction pathways in silico– a theme of which is the utilization of many different data sources integrating chemical, drug and free text as well as the connection of physical protein interaction pathways and gene expression pathways. There was even a project which dealt with the alignment of pathway graphs (topology).
Dealing with pathways especially by hand and in the form of a picture is probably the bane of many biologists’ existence– I think that the solutions we’ll see in the next few years will turn this task into simple data-in-data-out software components, much like the kind we have to deal with sequence alignments.
And now, back to the real world!
Addendum: My talk went very well 🙂
And here are my slides with a preview below.
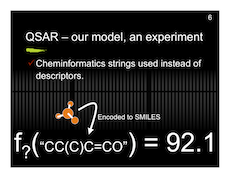


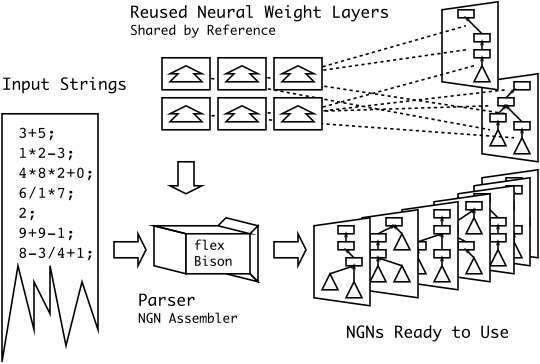
New Diagram for MSc-X3 (math paper)
Brief: I’m particularly happy with this diagram… I had something along these lines in my head for a while, but I never could figure out how to draw it correctly. I never thought that simplifying it to three easy steps was the smarter thing to do.
 Ed's Big Plans
Ed's Big Plans