Archive for the ‘Neural Networks’ tag
Blog Author Clustering with SOMs
For the final project in Neural Networks (CIS 6050 W11), I decided to cluster blog posts based on the difference between an author’s word choice and the word choice of the entire group of authors.
>>> Attached Course Paper: Kohonen Self-Organizing Maps in Clustering of Blog Authors (pdf) <<<
A self-organizing map (Kohonen SOM) strategy was used. The words chosen to compose a given blog post defined wherein the map it should be placed. The purpose of this project was to figure out what predictive power could be harnessed given the combination of the SOM and each author’s lexicon; i.e. whether or not it is possible to automatically categorize an author’s latest post without the use of any tools besides the above.
Data: Thirteen authors contributing a total of fourteen blogs participated in the study (all data was retrieved on 2011 March 8th). The below table summarizes the origin of the data.
| Author | Posts | Lexicon | Blog Name | Subject Matter |
| Andre Masella | 198 | 7953 | MasellaSphere | Synth Bio, Cooking, Engineering |
| Andrew Berry | 46 | 2630 | Andrew Berry Development | Drupal, Web Dev, Gadgets |
| Arianne Villa | 41 | 1217 | …mrfmr. | Internet Culture, Life |
| Cara Ma | 12 | 854 | Cara’s Awesome Blog | Life, Pets, Health |
| Daniela Mihalciuc | 211 | 4454 | Citisen of the World† | Travel, Life, Photographs |
| Eddie Ma | 161 | 5960 | Ed’s Big Plans | Computing, Academic, Science |
| Jason Ernst | 61 | 3445 | Jason’s Computer Science Blog | Computing, Academic |
| John Heil | 4 | 712 | Dos Margaritas Por Favor | Science, Music, Photography |
| Lauren Stein | 91 | 4784 | The Most Interesting Person | Improv, Happiness, Events |
| Lauren Stein (Cooking) | 7 | 593 | The Laurentina Cookbook | Cooking, Humour |
| Liv Monck-Whipp | 30 | 398 | omniology | Academic, Biology, Science |
| Matthew Gingerich | 98 | 395 | The Majugi Blog | Academic, Synth Bio, Engineering |
| Richard Schwarting | 238 | 7538 | Kosmokaryote | Academic, Computing, Linux |
| Tony Thompson | 51 | 2346 | Tony Thompson, Geek for Hire | Circuitry, Electronic Craft, Academic |
†Daniela remarks that the spelling of Citisen is intentional.
In order to place the blog posts into a SOM, each post was converted to a bitvector. Each bit is assigned to a specific word, so that the positions of each bit consistently represents the same word from post to post. An on-bit represented the presence of a word while an off-bit represented the absence of a word. Frequently used words like “the” were omitted from the word bit-vector, and seldom used words were also omitted.
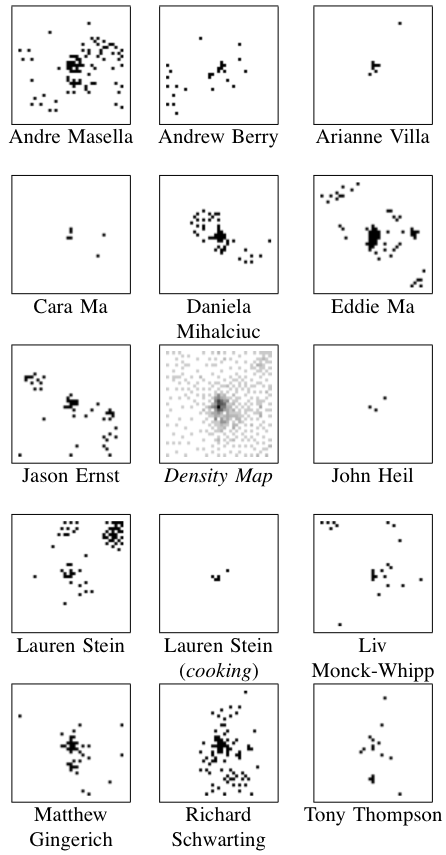 Results: The center image (in the collection to the left) is a density map where darker pixels indicates a larger number of posts — this centre map represents all of the posts made by all of the authors pooled together.
Results: The center image (in the collection to the left) is a density map where darker pixels indicates a larger number of posts — this centre map represents all of the posts made by all of the authors pooled together.
Because of the number of posts and the number of authors, I’ve exploded the single SOM image into the remaining fourteen images.
It was found that posts were most often clustered together if they were both by the same author and on the same topic. Clusters containing more than one author generally did not show much agreement about the topic.
Regions of this SOM were dominated by particular authors and topics as below.
| Region | Authors | Topics |
| Top Left | Liv | Academic Journals |
| Eddie | Software Projects | |
| Jason | Academic | |
| Top Border | Lauren | Human Idiosyncrasies |
| Richard | Linux | |
| Top Right | Lauren | Improv |
| Up & Left of Centre | Daniela | Travel |
| Centre | all | short and misfit posts |
| Right Border | Andre | Cooking |
| Just Below Centre | Matthew | Software Projects |
| Bottom Left | Andre | Language Theory |
| Andrew | Software Projects | |
| Jason | Software Projects | |
| Bottom Border | Richard | Academic |
| Bottom Right | Eddie | Web Development |
| Jason | Software Tutorials |
Discussion: There are some numerical results to go along with this, but they aren’t terribly exciting — the long and the short of it is that this project should to be repeated. The present work points towards the following two needed improvements.
First, the way the bitvectors were cropped at the beginning and at the end were based on a usage heuristic that doesn’t really conform to information theory. I’d likely take a look at the positional density of all bits to select meaningful words to cluster.
Second, all posts were included — this results in the dense spot in the middle of the central map. Whether these posts are short or just misfit, many of them can probably be removed by analyzing their bit density too.
Appendix: Here are two figures that describe the distribution of the data in the word bitvectors.
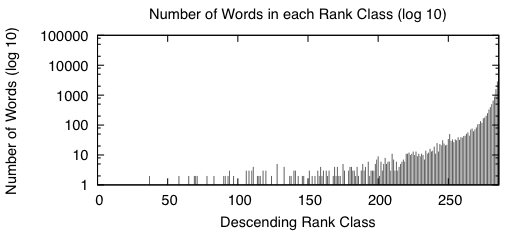 |
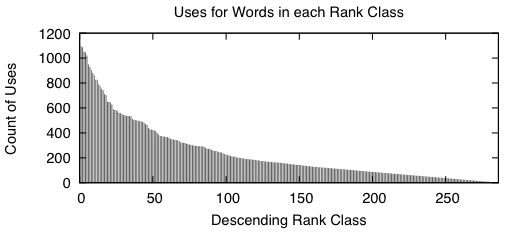 |
When we sort the words based from a high number of occurrences down to a low number of occurrences, we get graphs that look like the above two. A rank class contains all words that have the same number of occurrences across the entire study. The impulse graph on the left shows the trend for the number of unique words in each rank class. The number of words drastically increases as the classes contain fewer words. The impulse graph on the right shows the trend for the count of uses for words in a given rank class. The number of uses decreases as words become more rare.
These graphs were made before the main body of the work to sketch out how I wanted the bitvectors to behave — they verify that there was nothing unusual about the way the words were distributed amongst the data.
 Ed's Big Plans
Ed's Big Plans