TIM Barrels and 4-Alpha Helix Bundles
Beta-Alpha TIM Barrels and 4-Alpha Helix bundles are the first of the major folds I’ll be looking at here at Waterloo…
As with all academic projects, the probability of goal, approach and method mutation is high. Looking at the above protein folds serves as an excellent starting point as I’ll be applying some of the established methods that Andrew and Aaron have developed.
Alpha helices and Beta sheets are objects that any highschool biologist is acquainted with. To recapitulate, alpha helices are sequences of amino acids arranged so that the alpha carbon of each amino acid falls along the path of a helix. The number of amino acids per turn in this peptide and the regularity of the helix are determined by both the sequence and the environment that peptide finds itself in. Amino acids in the beta sheet conformation are arranged so that their alpha carbons zig zag. The result is a nice wide and flat shape schematically drawn as a sheet. Alpha helices and Beta sheets are collectively called secondary structural elements.
So, folds are these giant overarching classification of proteins– Folds themselves are inherently structural, so classifying them OR using them as classifications is only relevant in structural studies and databases on the web like SCOP and CATH. In databases such as SCOP and CATH, classification of similarly structured proteins start by determining whether the protein contains mostly alpha helices; mostly beta sheets; beta sheet and alpha helices alternatively and irregularly; or beta sheet and alpha helices in distinct regions of a protein. In SCOP, further classification is done by manually assigning proteins to smaller and smaller categories, while in CATH, these classifications are done by a hidden Markov model and then manually inspected (or not). It turns out that CATH uses a similar manual approach, and uses HMMs only to assist; contrast with PFAM which actually utilizes HMMs for the majority of work and is verified afterward by humans.
Certain folds like the beta-Trefoil and TIM Barrel benefit from containing only proteins that cleanly fit into some subcategory or several subcategories– it is then possible to just drill into the right level of categorization and pull out all of the beta-Trefoils and TIM Barrels we want.
The 4-Alpha-Helix Bundle constitutes a fold of protein that manages to be spread around the databases, being a very common secondary structural repeat; also a very small repeat when compared to the two giants above. These two items represent an interesting contrast too. Both machine and human intelligence pulls TIM Barrels together while sprinkling alpha-helix bundles across databases and subcategories. And yes, the size difference helps too.
So, I’m starting with a structural then sequence based alignment for single domain TIM Barrels and alpha-helix bundles; to be completely focused, an objective is named: To identify where sequence repeats occur in each individual protein.
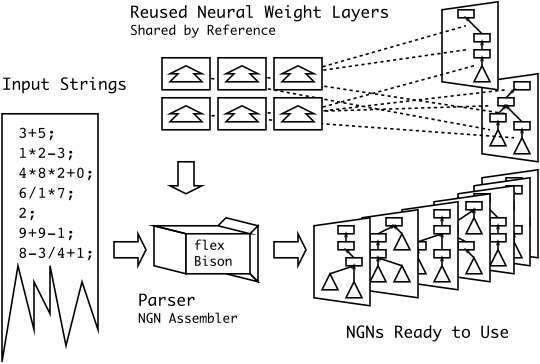
New Diagram for MSc-X3 (math paper)
Brief: I’m particularly happy with this diagram… I had something along these lines in my head for a while, but I never could figure out how to draw it correctly. I never thought that simplifying it to three easy steps was the smarter thing to do.
Monet Molecules c/o Autotrace
A Monet Isopentenol c/o Autotrace
Brief: Happy accidents make me happy– here’s isopentenol after grabbing it as an SDF out of PubChem, dumping it out into a PNG with Bioclipse, grayscaling it with GIMP— then converting it to an SVG with Autotrace (RO IT Systems)… It’s just pretty… like a Monet. Of course, I have to go back and make it look like a molecule again for a paper… but I’m going to admire the pretty little alcohol for a bit.
 Ed's Big Plans
Ed's Big Plans